Intraspecific variation in grass microbiome interaction
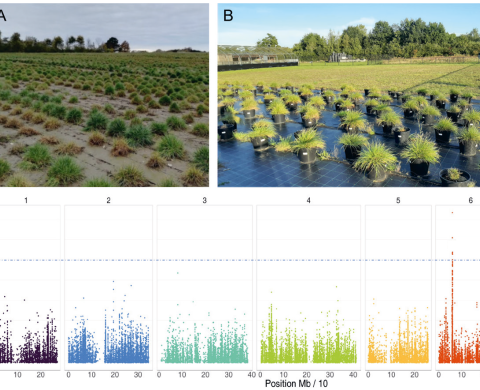
Through association with beneficial bacteria and fungi, plants can express increased growth via improved nutrient uptake, disease resistance, and abiotic stress tolerance. Such microbe-mediated plant traits could be important for ecological adaptation and crop improvement, but natural or artificial selection can only shape these traits if genetic variation exists for the ability to engage with beneficial microbes. While we know that plant species can significantly differ in their associated microbiomes and other soil biotas, we know little about how much intraspecific variation there is for these associations. Grasses are a good model for addressing this question of intraspecific variation. Indeed, grass species harbor extensive genetic diversity, which is reflected in their tolerance to different habitats and conditions, and therefore, they represent valuable material for future breeding programs. In this PhD project, I will use a broad panel of genotypes of Lolium perenne, Festuca arundinacea and Poa pratensis grass species to gain a better understanding of the genetic variation of microbiome recruitment and its functional role under environmental stress.
Experts
-
-
Paola Rallo
PhD Candidate , Terrestrial Ecology
-
Wim H. van der Putten
Researcher , Terrestrial Ecology
